Academic Staff and Fellows

- KAWASHITA Norihito
- Associate Professor Ph. D. in Pharmaceutical Sciences
-
Department/Energy and Materials
Department of Energy and Materials
Graduate school/Science Innovative Engineering(兼担)
We aim to elucidate various phenomena in the life sciences, including molecular evolution, drug susceptibility, and protein-protein interactions, using computational chemistry and bioinformatics that utilize computers and supercomputer.
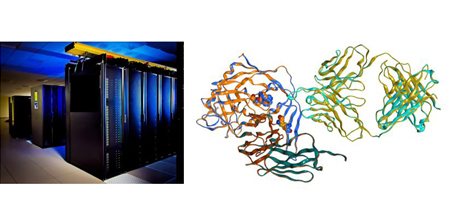
Life Science with Computer
| Research Area | Computational Chemistry, Bioinformatics, Machine Learning, Protein-protein interaction, Infections, Molecular Design, Drug Resistance, Molecular Evolution |
|---|---|
| Teaching (Undergraduate Course) | <Department of Energy and Materials> Introduction to Energy and Materials Exercises for Mathematics in Chemistry Exercises for Mathematics in Physics Synthetic Seminar 1 and 2 <Department of Life Science> Seminar for Life Science 1 and 2 Individual Study for Bachelor Thesis |
| Teaching (Graduate Course) | Computational Life Science Introduction to Advanced Studies on Biomolecules and Environmental Chemistry Advanced Studies on Biomolecules and Environmental Chemistry Bioinformatics for Genetic Counseling (Genetic Counseling Course) Advanced Exercise in Human Genetics (Genetic Counseling Course) Clinical Genetics 1 (Genetic Counseling Course) Genetic Counseling II (Genetic Counseling Course) Supplemental Nutrition Substances (Innovative Engineering) Exercise Lesson in Innovative Engineering (Innovative Engineering) Master Study in Innovative Engineering (Innovative Engineering) |
| Research Interests | Protein-protein interaction analysis by supercomputer "Fugaku". Drug design related studies for protein-protein interaction inhibitor Identification of pathogenic factor and mechanism of mutation by molecular evolution method Identification of hereditary diseases and virus infection mechanism by molecular modeling |
| Selected Publications |
(1) Kawashita N, Interaction Analysis by Fragment Molecular Orbital Method for Drug Discovery Research. Chem. Pharm. Bull., in press. (2) Takaba K, Watanabe C, Tokuhisa A, Akinaga Y, Ma B, Kanada R, Araki M, Okuno Y, Kawashima Y, Moriwaki H, Kawashita N, Honma T, Fukuzawa K, Tanaka S. Protein-ligand binding affinity prediction of cyclin-dependent kinase-2 inhibitors by dynamically averaged fragment molecular orbital-based interaction energy. J. Comput. Chem., 43(20), 1362-1371, (2022). doi: 10.1002/jcc.26940 (3) Watanabe K, Watanabe C, Honma T, Tian YS, Kawashima Y, Kawashita N, Takagi T, Fukuzawa K. Intermolecular Interaction Analyses on SARS-CoV-2 Spike Protein Receptor Binding Domain and Human Angiotensin-Converting Enzyme 2 Receptor-Blocking Antibody/Peptide Using Fragment Molecular Orbital Calculation. J. Phys. Chem. Lett., 12(16), 4059-4066, (2021). |
| Research and Achievements |
See Resrarchmap.
|
| Education (Undergraduate Course) |
Department of Material Science, Faculty of Science, Osaka City University |
| Education (Master's/Doctral Course) |
Graduate School of Pharmaceutical Sciences, Osaka University |
| Biography | Designated Research Associate, Designated Assistant Professor, Research Institute for Microbial Diseases, Osaka University (2005-2007) Assistant Professor, Graduate School of Pharmaceutical Sciences, Osaka University (2007-2017) Lecturer, Department of Life Science, Kindai University (2017-2022) Associate Professor, Department of Energy and Material, Kindai University (2022-present) |
Computational Life Science
| Office Location | 4th floor, 38th building, HigashiOsaka campus |
|---|---|
nkawashita(at)emat.kindai.ac.jp
|
|
| Laboratory URL |
https://www.life.kindai.ac.jp/laboratory/kawashita/ |